Protein Window
The protein window is a popup window that appears when you click on a particular protein. The protein window shows information about the protein and also provide links to other sections of STRING or to other external websites.
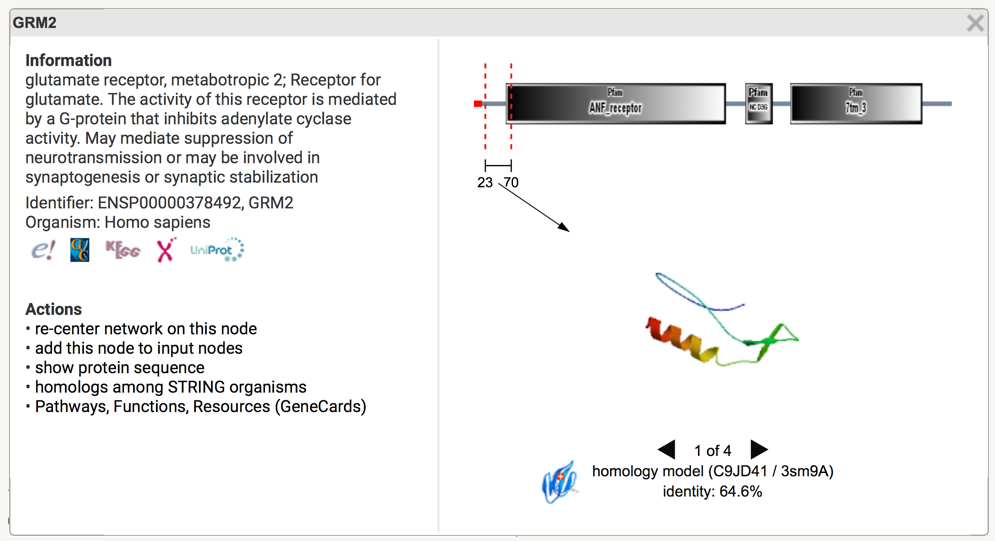
The protein window is divided in two part: left and right. The right part is displayed only when we have the protein structure (or a model of the protein structure) stored in our database. Note that for each new release of STRING we store in the images of crystallized protein (from PDB) as well as the images of protein models (from SwissModel). Since both PDB and SwissModel are updated more frequently than STRING, we suggest to directly look for structures/models on the PDB and SwissModel websites.
Information Top ↑
A description of the protein is shown. The identifier of the protein generally correspond to the identifiers of the import source of the genomic information (e.g. Ensembl) and the the gene name. The organism is indicated for clarity. Also links to other resources are provided (e.g., UniProt, Ensembl, etc.).
Actions Top ↑
The action are available are:
- recenter the network on this node - adjusts the network to this protein (i.e. look for all the proteins in STRING that interact with the protein of the window). This is a shortcut with respect to move back to the input page and relaunch STRING with the corresponding protein identifier
- add this node to the input node - adds to the network also the proteins that interact with the protein of the window. This is a shortcut with respect to moving back to the input page and relaunch STRING with both the identifiers (i.e. the input protein and the selected protein)
- show protein sequence - opens popup window containing the amino acid sequence of the protein
- homologs in STRING - brings to a String page where you can see all the homologs of your protein in the same and also in other organisms.
- Pathway, Functions, Resources (GeneCards) - redirects to the relative entry in the GeneCards database (only for human proteins).
- show protein domain (SMART) - redirects to the relative entry in the SMART database.
Domain and Structure Top ↑
Note that if no crystal-structure or structure model can be found in our database for the protein (or for similar proteins), this part is not shown.
In the upper part you will see colored shapes placed on a line. The shapes represent different SMART domains and the line represents the protein sequence. When no domains are found for the protein you just see a grey line with length proportional to the length of the protein sequence. When the protein is longer than 800 amino acids you will see some dots at the beginning (or end) of the line, indicating that the protein is too long to be displayed on the window. Clicking on the SMART image you will be redirected to the corresponding entry in the SMART database.
Below the SMART domains you will see an image of the protein structure. This can be either the image of a crystal structure (taken from PDB) or the model of the structure of the protein (taken from SwissModel). You can distinguish these two cases simply looking at the small logo just below the structure. Clicking on the structure you will be redirected to the corresponding entry in the PDB (or SwissModel) database.
Usually the image of the protein structure represents only a part of the protein (domain). To illustrate this, an arrow connects a section of the SMART image to the structure, indicating the region of the protein that is represented by the structure image. Sometimes a protein may contain more than one identical domain. In this case you will see several arrows connecting different parts of the protein to the structure.
For the PDB structures whole protein structure is displayed, when this is available. In this case, hovering the mouse cursor over the image will change the image of the whole protein structure with the image of the chain (i.e. the structure of only that part of the protein indicated by the arrow).
When more than a structure/model is present for a protein, two little arrows appear at the bottom. You can click on those arrows to move between the different structures.
The image of the structure may be that of a protein that is similar in sequence to your protein For both PDB and SwissModel we indicate the identity. The identity refers to the alignment of the STRING protein with the PDB protein. This alignment is the result of an all-against-all blast search that we execute for all the STRING proteins against all the PDB proteins (for the SwissModel's protein a similar alignment against PDB templates is performed by the swissModel group). For this reason, only an identity of 100% indicate that the displayed image is the structure of your protein (andnot of a similar one).